
Alternative splicing
process of generating multiple mRNA molecules from a given set of exons by differential use of exons from the primary transcript(s), to form multiple mature mRNAs / From Wikipedia, the free encyclopedia
Alternative splicing allows DNA to code for more than one protein. It varies the exon make-up of the messenger RNA.
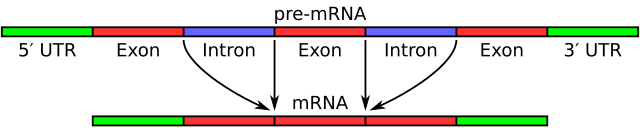

In alternative splicing the exons of the pre-messenger RNA produced by transcription are reconnected in different ways during RNA splicing.
This produces different mature messenger RNAs from the same gene. They get translated into different proteins. Thus, a single gene may code for multiple proteins.[1]
Alternative splicing is normal in eukaryotes. It greatly increases the diversity of proteins that can be encoded by the genome.[1] In humans, ~95% of multiexonic genes are alternatively spliced.[2][3][4]
There are various kinds of alternative splicing: the most common is exon skipping. An exon may be included in mRNAs under some conditions or in particular tissues, and omitted from the mRNA in others.[1] There are splicing activators that promote the use of a particular splice site, and splicing repressors that reduce the use of a particular site. New types of alternative splicing are being found.[4][5]
Abnormal variations in splicing occur in disease. Many human genetic disorders come from splicing variants.[4] Abnormal splicing variants may also contribute to the development of cancer.[6][7][8] High-throughput sequencing of RNA can for example be used to measure the genome-scale amount of deviating alternative splicing, such as in a cohort of colorectal cancers, where high amount of aberrant splicing was associated to poor patient survival.[9] Non-working splicing products are usually dealt with by post-transcriptional quality control.[10] That is, they are chopped up by enzymes.