利用者:Iaacc/sandbox
ウィキペディア フリーな encyclopedia
https://en.wikipedia.org/w/index.php?title=RNA-Seq&oldid=916228806
 |
ここはIaaccさんの利用者サンドボックスです。編集を試したり下書きを置いておいたりするための場所であり、百科事典の記事ではありません。ただし、公開の場ですので、許諾されていない文章の転載はご遠慮ください。
登録利用者は自分用の利用者サンドボックスを作成できます(サンドボックスを作成する、解説)。 その他のサンドボックス: 共用サンドボックス | モジュールサンドボックス 記事がある程度できあがったら、編集方針を確認して、新規ページを作成しましょう。 |
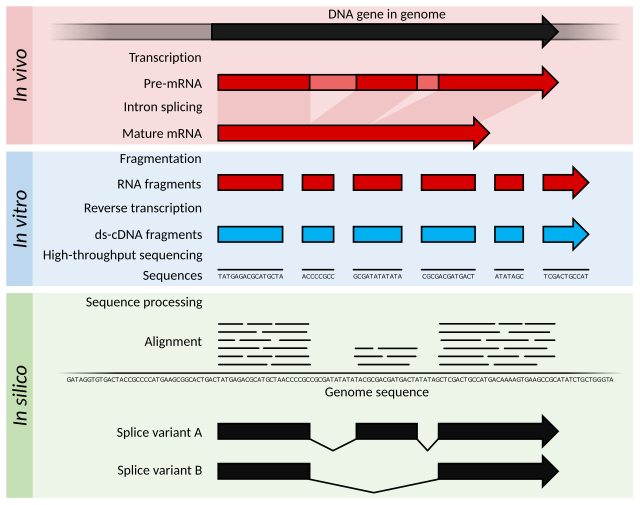
RNA-seqとは、次世代DNAシークエンシング(NGS)を用いて、生物試料中のRNAの検出や定量を行う解析のこと。
RNA-Seq uses next-generation sequencing (NGS) to reveal the presence and quantity of RNA in a biological sample at a given moment, analyzing the continuously changing cellular transcriptome.[2][3]
とりわけ、RNA-seqは
Specifically, RNA-Seq facilitates the ability to look at alternative gene spliced transcripts, post-transcriptional modifications, gene fusion, mutations/SNPs and changes in gene expression over time, or differences in gene expression in different groups or treatments.[4] In addition to mRNA transcripts, RNA-Seq can look at different populations of RNA to include total RNA, small RNA, such as miRNA, tRNA, and ribosomal profiling.[5] RNA-Seq can also be used to determine exon/intron boundaries and verify or amend previously annotated 5' and 3' gene boundaries. Recent advances in RNA-Seq include single cell sequencing and in situ sequencing of fixed tissue.[6]
Prior to RNA-Seq, gene expression studies were done with hybridization-based microarrays. Issues with microarrays include cross-hybridization artifacts, poor quantification of lowly and highly expressed genes, and needing to know the sequence a priori.[7] Because of these technical issues, transcriptomics transitioned to sequencing-based methods. These progressed from Sanger sequencing of Expressed Sequence Tag libraries, to chemical tag-based methods (e.g., serial analysis of gene expression), and finally to the current technology, next-gen sequencing of cDNA (notably RNA-Seq).