Loading AI tools
Class of enzymes From Wikipedia, the free encyclopedia
Tryptophan synthase or tryptophan synthetase is an enzyme (EC 4.2.1.20) that catalyzes the final two steps in the biosynthesis of tryptophan.[1][2] It is commonly found in Eubacteria,[3] Archaebacteria,[4] Protista,[5] Fungi,[6] and Plantae.[7] However, it is absent from Animalia.[8] It is typically found as an α2β2 tetramer.[9][10] The α subunits catalyze the reversible formation of indole and glyceraldehyde-3-phosphate (G3P) from indole-3-glycerol phosphate (IGP). The β subunits catalyze the irreversible condensation of indole and serine to form tryptophan in a pyridoxal phosphate (PLP) dependent reaction. Each α active site is connected to a β active site by a 25 Ångstrom long hydrophobic channel contained within the enzyme. This facilitates the diffusion of indole formed at α active sites directly to β active sites in a process known as substrate channeling.[11] The active sites of tryptophan synthase are allosterically coupled.[12]
| Tryptophan Synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
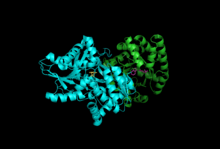 Subunits: Beta Subunit, Alpha Subunit with PLP, IGP | |||||||||
| Identifiers | |||||||||
| EC no. | 4.2.1.20 | ||||||||
| CAS no. | 9014-52-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||

Tryptophan synthase typically exists as an α-ββ-α complex. The α and β subunits have molecular masses of 27 and 43 kDa respectively. The α subunit has a TIM barrel conformation. The β subunit has a fold type II conformation and a binding site adjacent to the active site for monovalent cations.[13] Their assembly into a complex leads to structural changes in both subunits resulting in reciprocal activation. There are two main mechanisms for intersubunit communication. First, the COMM domain of the β-subunit and the α-loop2 of the α-subunit interact. Additionally, there are interactions between the αGly181 and βSer178 residues.[14] The active sites are regulated allosterically and undergo transitions between open, inactive, and closed, active, states.[12]
The α and β active sites are separated by a 25 Ångstrom long hydrophobic channel contained within the enzyme allowing for the diffusion of indole. If the channel did not exist, the indole formed at an α active site would quickly diffuse away and be lost to the cell as it is hydrophobic and can easily cross membranes. As such, the channel is essential for enzyme complex function.[15]

The net reaction of tryptophan synthase turns indole-3-glycerol phosphate and serine into glyceraldehyde-3-phosphate, tryptophan and water. The reaction happens in two steps, each catalyzed by one of the subunits:

The α subunit catalyzes the formation of indole and G3P from a retro-aldol cleavage of IGP. The αGlu49 and αAsp60 are thought to be directly involved in the catalysis as shown.[11] The rate limiting step is the isomerization of IGP.[16] See image 2.
The β subunit catalyzes the β-replacement reaction in which indole and serine condense to form tryptophan in a PLP dependent reaction. The βLys87, βGlu109, and βSer377 are thought to be directly involved in the catalysis as shown.[11] Again, the exact mechanism has not been conclusively determined. See image 2.
Tryptophan synthase is commonly found in Eubacteria, Archaebacteria, Protista, Fungi, and Plantae. It is absent from animals such as humans. Tryptophan is one of the twenty standard amino acids and one of nine essential amino acids for humans. As such, tryptophan is a necessary component of the human diet.
Tryptophan synthetase is also known to accept indole analogues, e.g., fluorinated or methylated indoles, as substrates, generating the corresponding tryptophan analogues.[17]
As humans do not have tryptophan synthase, this enzyme has been explored as a potential drug target.[18] However, it is thought that bacteria have alternate mechanisms to produce amino acids which might make this approach less effective. In either case, even if the drug only weakens bacteria, it might still be useful as the bacteria are already vulnerable in the hostile host environment. As such, the inhibition of tryptophan synthase along with other PLP-enzymes in amino acid metabolism has the potential to help solve medical problems.[19]
Inhibition of tryptophan synthase and other PLP-enzymes in amino acid metabolism has been suggested for:
It is thought that early in evolution the trpB2 gene was duplicated. One copy entered the trp operon as trpB2i allowing for its expression with trpA. TrpB2i formed transient complexes with TrpA and in the process activated TrpA unidirectionally. The other copy remained outside as trpB2o, and fulfilled an existing role or played a new one such as acting as a salvage protein for indole. TrpB2i evolved into TrpB1, which formed permanent complexes with trpA resulting in bidirectional activation. The advantage of the indole salvage protein declined and the TrpB gene was lost. Finally, the TrpB1 and TrpA genes were fused resulting in the formation the bifunctional enzyme.[22]
Tryptophan synthase was the first enzyme identified that had two catalytic capabilities that were extensively studied. It was also the first identified to utilize substrate channeling. As such, this enzyme has been studied extensively and is the subject of great interest.[11]
Seamless Wikipedia browsing. On steroids.
Every time you click a link to Wikipedia, Wiktionary or Wikiquote in your browser's search results, it will show the modern Wikiwand interface.
Wikiwand extension is a five stars, simple, with minimum permission required to keep your browsing private, safe and transparent.