Loading AI tools
Cellular appendage functioning as locomotive or sensory organelle From Wikipedia, the free encyclopedia
A flagellum (/fləˈdʒɛləm/; pl.: flagella) (Latin for 'whip' or 'scourge') is a hair-like appendage that protrudes from certain plant and animal sperm cells, from fungal spores (zoospores), and from a wide range of microorganisms to provide motility.[1][2][3][4] Many protists with flagella are known as flagellates.
| Flagellum | |
|---|---|
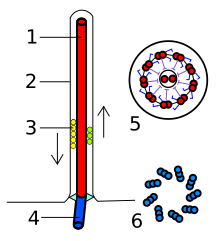
 Structure of bacterial flagellum | |
 | |
| Identifiers | |
| MeSH | D005407 |
| TH | H1.00.01.1.01032 |
| FMA | 67472 |
| Anatomical terminology | |
A microorganism may have from one to many flagella. A gram-negative bacterium Helicobacter pylori, for example, uses its flagella to propel itself through the stomach to reach the mucous lining where it may colonise the epithelium and potentially cause gastritis, and ulcers – a risk factor for stomach cancer.[5] In some swarming bacteria, the flagellum can also function as a sensory organelle, being sensitive to wetness outside the cell.[6]
Across the three domains of Bacteria, Archaea, and Eukaryota, the flagellum has a different structure, protein composition, and mechanism of propulsion but shares the same function of providing motility. The Latin word flagellum means "whip" to describe its lash-like swimming motion. The flagellum in archaea is called the archaellum to note its difference from the bacterial flagellum.[7][8]
Eukaryotic flagella and cilia are identical in structure but have different lengths and functions.[9] Prokaryotic fimbriae and pili are smaller, and thinner appendages, with different functions. Cilia are attached to the surface of flagella and are used to swim or move fluid from one region to another. [10]

The three types of flagella are bacterial, archaeal, and eukaryotic.
The flagella in eukaryotes have dynein and microtubules that move with a bending mechanism. Bacteria and archaea do not have dynein or microtubules in their flagella, and they move using a rotary mechanism.[12]
Other differences among these three types are:
The bacterial flagellum is made up of protein subunits of flagellin.[12] Its shape is a 20-nanometer-thick hollow tube. It is helical and has a sharp bend just outside the outer membrane; this "hook" allows the axis of the helix to point directly away from the cell. A shaft runs between the hook and the basal body, passing through protein rings in the cell's membrane that act as bearings. Gram-positive organisms have two of these basal body rings, one in the peptidoglycan layer and one in the plasma membrane. Gram-negative organisms have four such rings: the L ring associates with the lipopolysaccharides, the P ring associates with peptidoglycan layer, the M ring is embedded in the plasma membrane, and the S ring is directly attached to the cytoplasm. The filament ends with a capping protein.[21][22]
The flagellar filament is the long, helical screw that propels the bacterium when rotated by the motor, through the hook. In most bacteria that have been studied, including the gram-negative Escherichia coli, Salmonella typhimurium, Caulobacter crescentus, and Vibrio alginolyticus, the filament is made up of 11 protofilaments approximately parallel to the filament axis. Each protofilament is a series of tandem protein chains. However, Campylobacter jejuni has seven protofilaments.[23]
The basal body has several traits in common with some types of secretory pores, such as the hollow, rod-like "plug" in their centers extending out through the plasma membrane. The similarities between bacterial flagella and bacterial secretory system structures and proteins provide scientific evidence supporting the theory that bacterial flagella evolved from the type-three secretion system (TTSS).
The atomic structure of both bacterial flagella as well as the TTSS injectisome have been elucidated in great detail, especially with the development of cryo-electron microscopy. The best understood parts are the parts between the inner and outer membrane, that is, the scaffolding rings of the inner membrane (IM), the scaffolding pairs of the outer membrane (OM), and the rod/needle (injectisome) or rod/hook (flagellum) sections.[24]

The bacterial flagellum is driven by a rotary engine (Mot complex) made up of protein, located at the flagellum's anchor point on the inner cell membrane. The engine is powered by proton-motive force, i.e., by the flow of protons (hydrogen ions) across the bacterial cell membrane due to a concentration gradient set up by the cell's metabolism (Vibrio species have two kinds of flagella, lateral and polar, and some are driven by a sodium ion pump rather than a proton pump[26]). The rotor transports protons across the membrane, and is turned in the process. The rotor alone can operate at 6,000 to 100,000 rpm,[27] but with the flagellar filament attached usually only reaches 200 to 1000 rpm. The direction of rotation can be changed by the flagellar motor switch almost instantaneously, caused by a slight change in the position of a protein, FliG, in the rotor.[28] The torque is transferred from the MotAB to the torque helix on FliG's D5 domain and with the increase in the requirement of the torque or speed more MotAB are employed.[25] Because the flagellar motor has no on-off switch, the protein epsE is used as a mechanical clutch to disengage the motor from the rotor, thus stopping the flagellum and allowing the bacterium to remain in one place.[29]
The production and rotation of a flagellum can take up to 10% of an Escherichia coli cell's energy budget and has been described as an "energy-guzzling machine" [30]. Its operation generates reactive oxygen species that elevates mutation rates.[30]
The cylindrical shape of flagella is suited to locomotion of microscopic organisms; these organisms operate at a low Reynolds number, where the viscosity of the surrounding water is much more important than its mass or inertia.[31]
The rotational speed of flagella varies in response to the intensity of the proton-motive force, thereby permitting certain forms of speed control, and also permitting some types of bacteria to attain remarkable speeds in proportion to their size; some achieve roughly 60 cell lengths per second. At such a speed, a bacterium would take about 245 days to cover 1 km; although that may seem slow, the perspective changes when the concept of scale is introduced. In comparison to macroscopic life forms, it is very fast indeed when expressed in terms of number of body lengths per second. A cheetah, for example, only achieves about 25 body lengths per second.[32]
Through use of their flagella, bacteria are able to move rapidly towards attractants and away from repellents, by means of a biased random walk, with runs and tumbles brought about by rotating its flagellum counterclockwise and clockwise, respectively. The two directions of rotation are not identical (with respect to flagellum movement) and are selected by a molecular switch.[33] Clockwise rotation is called the traction mode with the body following the flagella. Counterclockwise rotation is called the thruster mode with the flagella lagging behind the body.[34]
During flagellar assembly, components of the flagellum pass through the hollow cores of the basal body and the nascent filament. During assembly, protein components are added at the flagellar tip rather than at the base.[35] In vitro, flagellar filaments assemble spontaneously in a solution containing purified flagellin as the sole protein.[36]
At least 10 protein components of the bacterial flagellum share homologous proteins with the type three secretion system (T3SS) found in many gram-negative bacteria,[37] hence one likely evolved from the other. Because the T3SS has a similar number of components as a flagellar apparatus (about 25 proteins), which one evolved first is difficult to determine. However, the flagellar system appears to involve more proteins overall, including various regulators and chaperones, hence it has been argued that flagella evolved from a T3SS. However, it has also been suggested[38] that the flagellum may have evolved first or the two structures evolved in parallel. Early single-cell organisms' need for motility (mobility) support that the more mobile flagella would be selected by evolution first,[38] but the T3SS evolving from the flagellum can be seen as 'reductive evolution', and receives no topological support from the phylogenetic trees.[39] The hypothesis that the two structures evolved separately from a common ancestor accounts for the protein similarities between the two structures, as well as their functional diversity.[40]
Some authors have argued that flagella cannot have evolved, assuming that they can only function properly when all proteins are in place. In other words, the flagellar apparatus is "irreducibly complex".[41] However, many proteins can be deleted or mutated and the flagellum still works, though sometimes at reduced efficiency.[42] Moreover, with many proteins unique to some number across species, diversity of bacterial flagella composition was higher than expected.[43] Hence, the flagellar apparatus is clearly very flexible in evolutionary terms and perfectly able to lose or gain protein components. For instance, a number of mutations have been found that increase the motility of E. coli.[44] Additional evidence for the evolution of bacterial flagella includes the existence of vestigial flagella, intermediate forms of flagella and patterns of similarities among flagellar protein sequences, including the observation that almost all of the core flagellar proteins have known homologies with non-flagellar proteins.[37] Furthermore, several processes have been identified as playing important roles in flagellar evolution, including self-assembly of simple repeating subunits, gene duplication with subsequent divergence, recruitment of elements from other systems ('molecular bricolage') and recombination.[45]
Different species of bacteria have different numbers and arrangements of flagella,[46][47] named using the term tricho, from the Greek trichos meaning hair.[48]
Counterclockwise rotation of a monotrichous polar flagellum pushes the cell forward with the flagellum trailing behind, much like a corkscrew moving inside cork. Water on the microscopic scale is highly viscous, unlike usual water.
Spirochetes, in contrast, have flagella called endoflagella arising from opposite poles of the cell, and are located within the periplasmic space as shown by breaking the outer-membrane and also by electron cryotomography microscopy.[51] The rotation of the filaments relative to the cell body causes the entire bacterium to move forward in a corkscrew-like motion, even through material viscous enough to prevent the passage of normally flagellated bacteria.
In certain large forms of Selenomonas, more than 30 individual flagella are organized outside the cell body, helically twining about each other to form a thick structure (easily visible with the light microscope) called a "fascicle".
In some Vibrio spp. (particularly Vibrio parahaemolyticus[52]) and related bacteria such as Aeromonas, two flagellar systems co-exist, using different sets of genes and different ion gradients for energy. The polar flagella are constitutively expressed and provide motility in bulk fluid, while the lateral flagella are expressed when the polar flagella meet too much resistance to turn.[53][54][55][56][57][58] These provide swarming motility on surfaces or in viscous fluids.
Bundling is an event that can happen in multi-flagellated cells, bundling the flagella together and causing them to rotate in a coordinated manner.
Flagella are left-handed helices, and when rotated counter-clockwise by their rotors, they can bundle and rotate together. When the rotors reverse direction, thus rotating clockwise, the flagellum unwinds from the bundle. This may cause the cell to stop its forward motion and instead start twitching in place, referred to as tumbling. Tumbling results in a stochastic reorientation of the cell, causing it to change the direction of its forward swimming.
It is not known which stimuli drive the switch between bundling and tumbling, but the motor is highly adaptive to different signals. In the model describing chemotaxis ("movement on purpose") the clockwise rotation of a flagellum is suppressed by chemical compounds favorable to the cell (e.g. food). When moving in a favorable direction, the concentration of such chemical attractants increases and therefore tumbles are continually suppressed, allowing forward motion; likewise, when the cell's direction of motion is unfavorable (e.g., away from a chemical attractant), tumbles are no longer suppressed and occur much more often, with the chance that the cell will be thus reoriented in the correct direction.
Even if all flagella would rotate clockwise, however, they often cannot form a bundle due to geometrical and hydrodynamic reasons.[59][60]



Aiming to emphasize the distinction between the bacterial flagella and the eukaryotic cilia and flagella, some authors attempted to replace the name of these two eukaryotic structures with "undulipodia" (e.g., all papers by Margulis since the 1970s)[61] or "cilia" for both (e.g., Hülsmann, 1992;[62] Adl et al., 2012;[63] most papers of Cavalier-Smith), preserving "flagella" for the bacterial structure. However, the discriminative usage of the terms "cilia" and "flagella" for eukaryotes adopted in this article (see § Flagella versus cilia below) is still common (e.g., Andersen et al., 1991;[64] Leadbeater et al., 2000).[65]
The core of a eukaryotic flagellum, known as the axoneme is a bundle of nine fused pairs of microtubules known as doublets surrounding two central single microtubules (singlets). This 9+2 axoneme is characteristic of the eukaryotic flagellum. At the base of a eukaryotic flagellum is a basal body, "blepharoplast" or kinetosome, which is the microtubule organizing center for flagellar microtubules and is about 500 nanometers long. Basal bodies are structurally identical to centrioles. The flagellum is encased within the cell's plasma membrane, so that the interior of the flagellum is accessible to the cell's cytoplasm.
Besides the axoneme and basal body, relatively constant in morphology, other internal structures of the flagellar apparatus are the transition zone (where the axoneme and basal body meet) and the root system (microtubular or fibrilar structures that extend from the basal bodies into the cytoplasm), more variable and useful as indicators of phylogenetic relationships of eukaryotes. Other structures, more uncommon, are the paraflagellar (or paraxial, paraxonemal) rod, the R fiber, and the S fiber.[66]: 63–84 For surface structures, see below.
Each of the outer 9 doublet microtubules extends a pair of dynein arms (an "inner" and an "outer" arm) to the adjacent microtubule; these produce force through ATP hydrolysis. The flagellar axoneme also contains radial spokes, polypeptide complexes extending from each of the outer nine microtubule doublets towards the central pair, with the "head" of the spoke facing inwards. The radial spoke is thought to be involved in the regulation of flagellar motion, although its exact function and method of action are not yet understood.[67]

The regular beat patterns of eukaryotic cilia and flagella generate motion on a cellular level. Examples range from the propulsion of single cells such as the swimming of spermatozoa to the transport of fluid along a stationary layer of cells such as in the respiratory tract.[68]
Although eukaryotic cilia and flagella are ultimately the same, they are sometimes classed by their pattern of movement, a tradition from before their structures have been known. In the case of flagella, the motion is often planar and wave-like, whereas the motile cilia often perform a more complicated three-dimensional motion with a power and recovery stroke.[68] Yet another traditional form of distinction is by the number of 9+2 organelles on the cell.[67]
Intraflagellar transport, the process by which axonemal subunits, transmembrane receptors, and other proteins are moved up and down the length of the flagellum, is essential for proper functioning of the flagellum, in both motility and signal transduction.[69]
Eukaryotic flagella or cilia, probably an ancestral characteristic,[70] are widespread in almost all groups of eukaryotes, as a relatively perennial condition, or as a flagellated life cycle stage (e.g., zoids, gametes, zoospores, which may be produced continually or not).[71][72][63]
The first situation is found either in specialized cells of multicellular organisms (e.g., the choanocytes of sponges, or the ciliated epithelia of metazoans), as in ciliates and many eukaryotes with a "flagellate condition" (or "monadoid level of organization", see Flagellata, an artificial group).
Flagellated lifecycle stages are found in many groups, e.g., many green algae (zoospores and male gametes), bryophytes (male gametes), pteridophytes (male gametes), some gymnosperms (cycads and Ginkgo, as male gametes), centric diatoms (male gametes), brown algae (zoospores and gametes), oomycetes (assexual zoospores and gametes), hyphochytrids (zoospores), labyrinthulomycetes (zoospores), some apicomplexans (gametes), some radiolarians (probably gametes),[73] foraminiferans (gametes), plasmodiophoromycetes (zoospores and gametes), myxogastrids (zoospores), metazoans (male gametes), and chytrid fungi (zoospores and gametes).
Flagella or cilia are completely absent in some groups, probably due to a loss rather than being a primitive condition. The loss of cilia occurred in red algae, some green algae (Zygnematophyceae), the gymnosperms except cycads and Ginkgo, angiosperms, pennate diatoms, some apicomplexans, some amoebozoans, in the sperm of some metazoans,[74] and in fungi (except chytrids).
A number of terms related to flagella or cilia are used to characterize eukaryotes.[72][75][66]: 60–63 [76][77] According to surface structures present, flagella may be:
According to the number of flagella, cells may be: (remembering that some authors use "ciliated" instead of "flagellated")[63][80]
According to the place of insertion of the flagella:[81]
According to the beating pattern:
Other terms related to the flagellar type:
The archaellum possessed by some species of Archaea is superficially similar to the bacterial flagellum; in the 1980s, they were thought to be homologous on the basis of gross morphology and behavior.[84] Both flagella and archaella consist of filaments extending outside the cell, and rotate to propel the cell. Archaeal flagella have a unique structure which lacks a central channel. Similar to bacterial type IV pilins, the archaeal proteins (archaellins) are made with class 3 signal peptides and they are processed by a type IV prepilin peptidase-like enzyme. The archaellins are typically modified by the addition of N-linked glycans which are necessary for proper assembly or function.[3]
Discoveries in the 1990s revealed numerous detailed differences between the archaeal and bacterial flagella. These include:
These differences support the theory that the bacterial flagella and archaella are a classic case of biological analogy, or convergent evolution, rather than homology.[87][88][89] Research into the structure of archaella made significant progress beginning in the early 2010s, with the first atomic resolution structure of an archaella protein, the discovery of additional functions of archaella, and the first reports of archaella in Nanoarchaeota and Thaumarchaeota.[90][91]
The only fungi to have a single flagellum on their spores are the chytrids. In Batrachochytrium dendrobatidis the flagellum is 19–20 μm long.[92] A nonfunctioning centriole lies adjacent to the kinetosome. Nine interconnected props attach the kinetosome to the plasmalemma, and a terminal plate is present in the transitional zone. An inner ring-like structure attached to the tubules of the flagellar doublets within the transitional zone has been observed in transverse section.[92]
Seamless Wikipedia browsing. On steroids.
Every time you click a link to Wikipedia, Wiktionary or Wikiquote in your browser's search results, it will show the modern Wikiwand interface.
Wikiwand extension is a five stars, simple, with minimum permission required to keep your browsing private, safe and transparent.